environmental DNA
monitoring biodiversity across Maryland
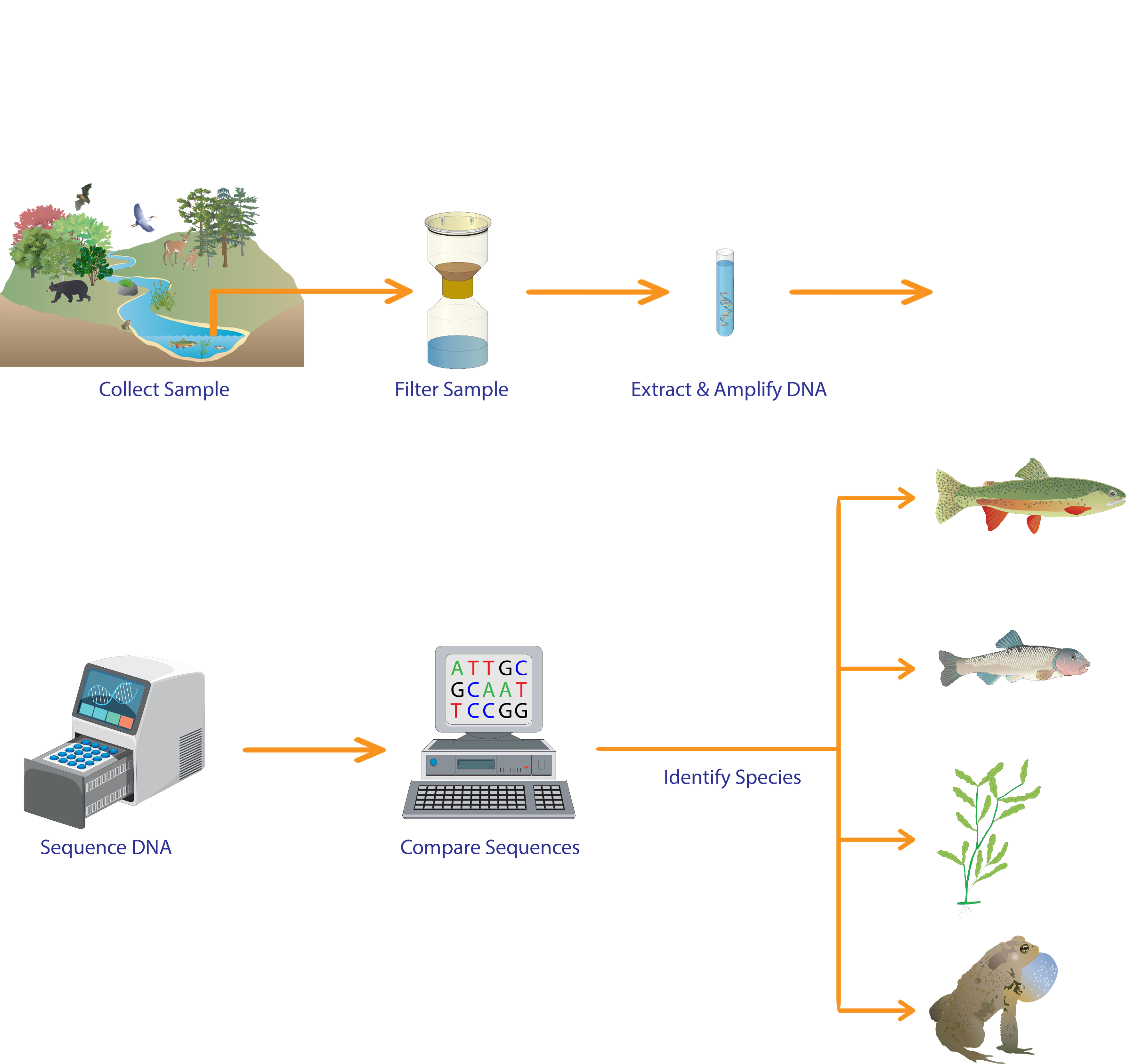
Environmental DNA (eDNA) allows detection of aquatic organisms from water samples alone—no netting, electrofishing, or visual surveys required. Aquatic organisms leave behind genetic traces that we collect and sequence to identify which species are present.
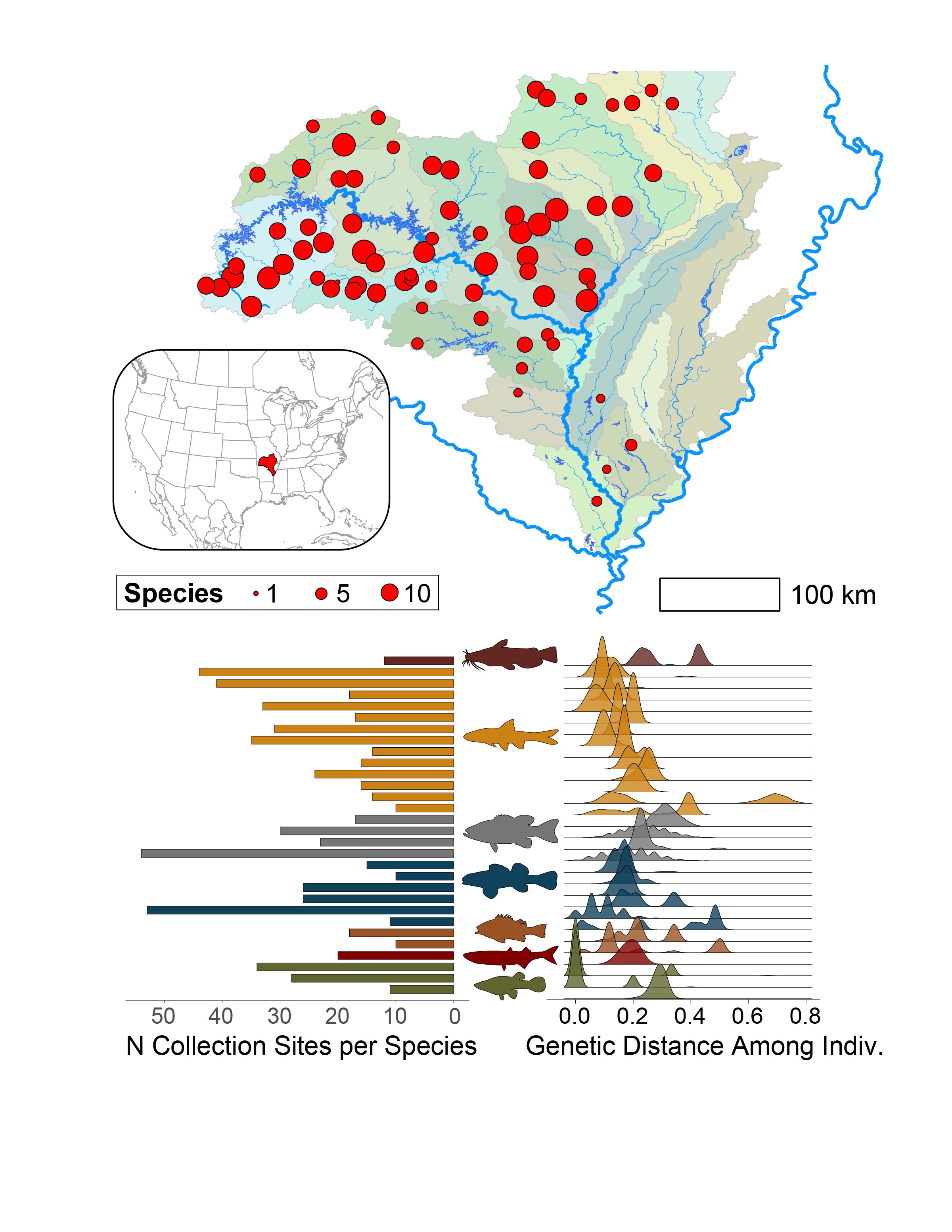
In collaboration with the Maryland Department of Natural Resources and the Maryland Biological Stream Survey (MBSS), we use eDNA to monitor biodiversity across Maryland's aquatic habitats, track rare and invasive species, and evaluate eDNA methods against conventional sampling.
Check out my publications page for updates on eDNA research.
In collaboration with the Maryland Department of Natural Resources and the Maryland Biological Stream Survey (MBSS), we use eDNA to monitor biodiversity across Maryland's aquatic habitats, track rare and invasive species, and evaluate eDNA methods against conventional sampling.
Check out my publications page for updates on eDNA research.